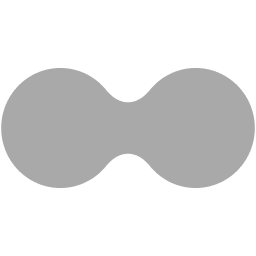
Overview of the order form
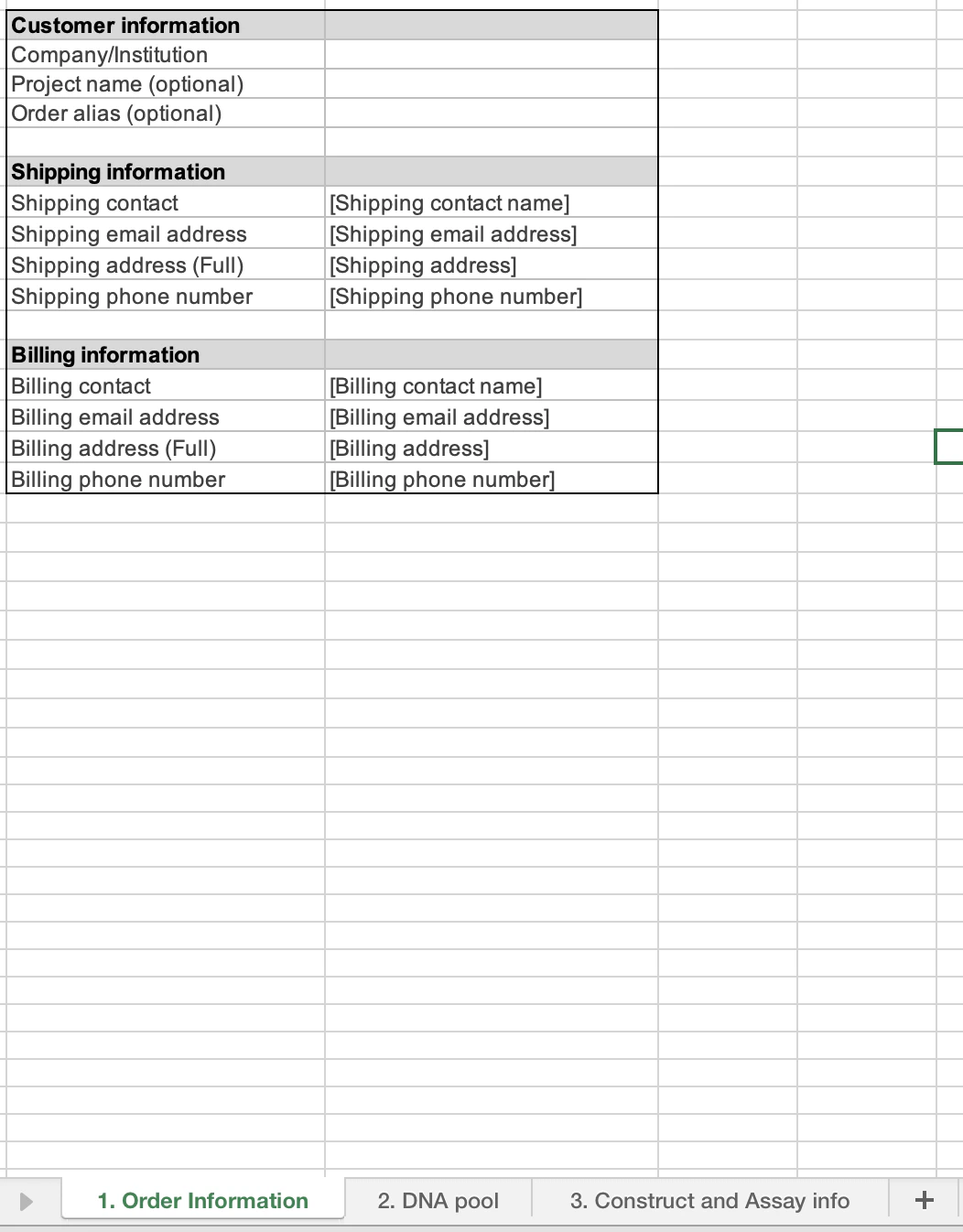
The order form consists of three tabs:- The Order information tab (1) requests general information about your order.
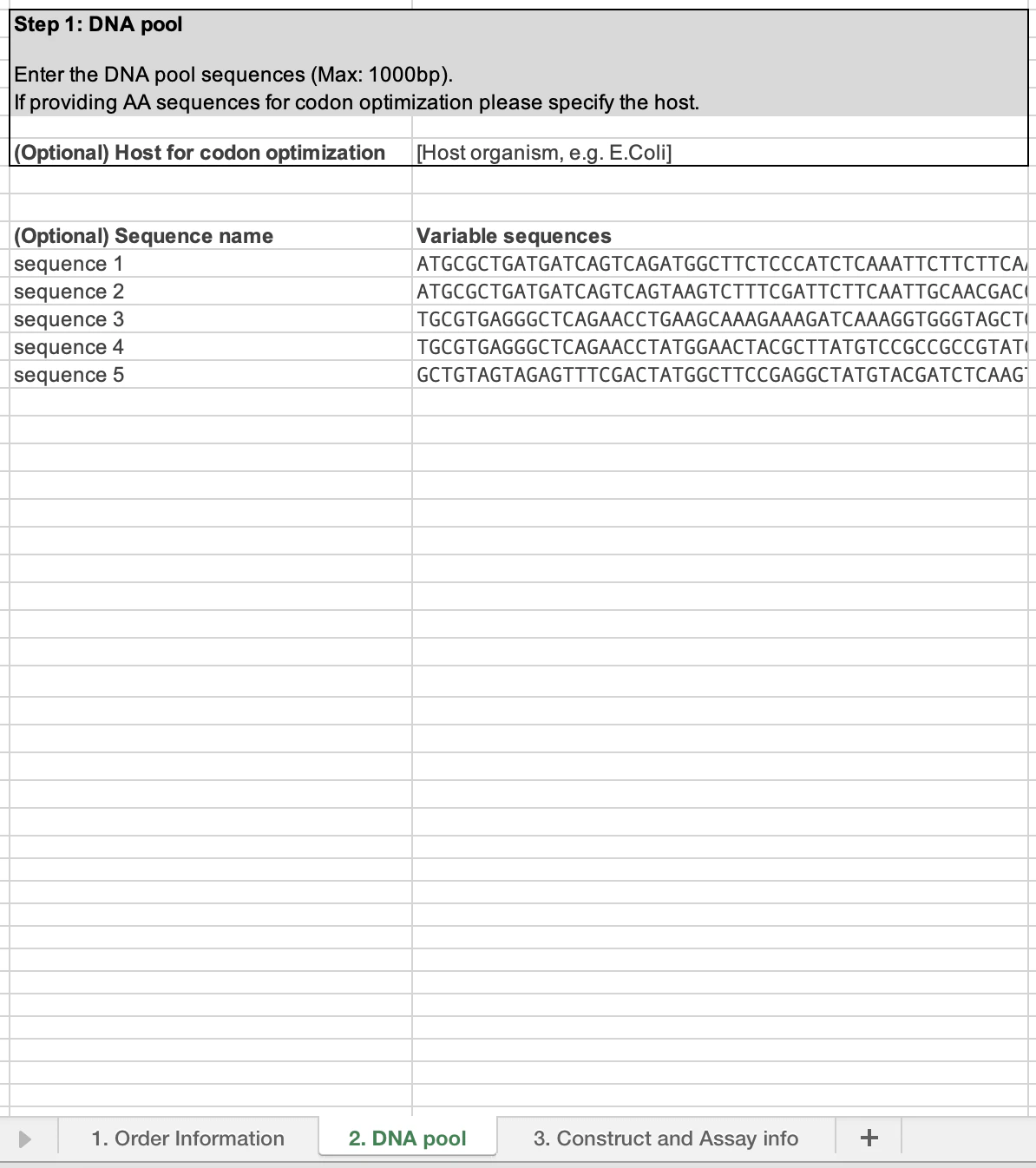
- The DNA pool tab (2) is where you can input the pool sequences you want to test and gain data on.
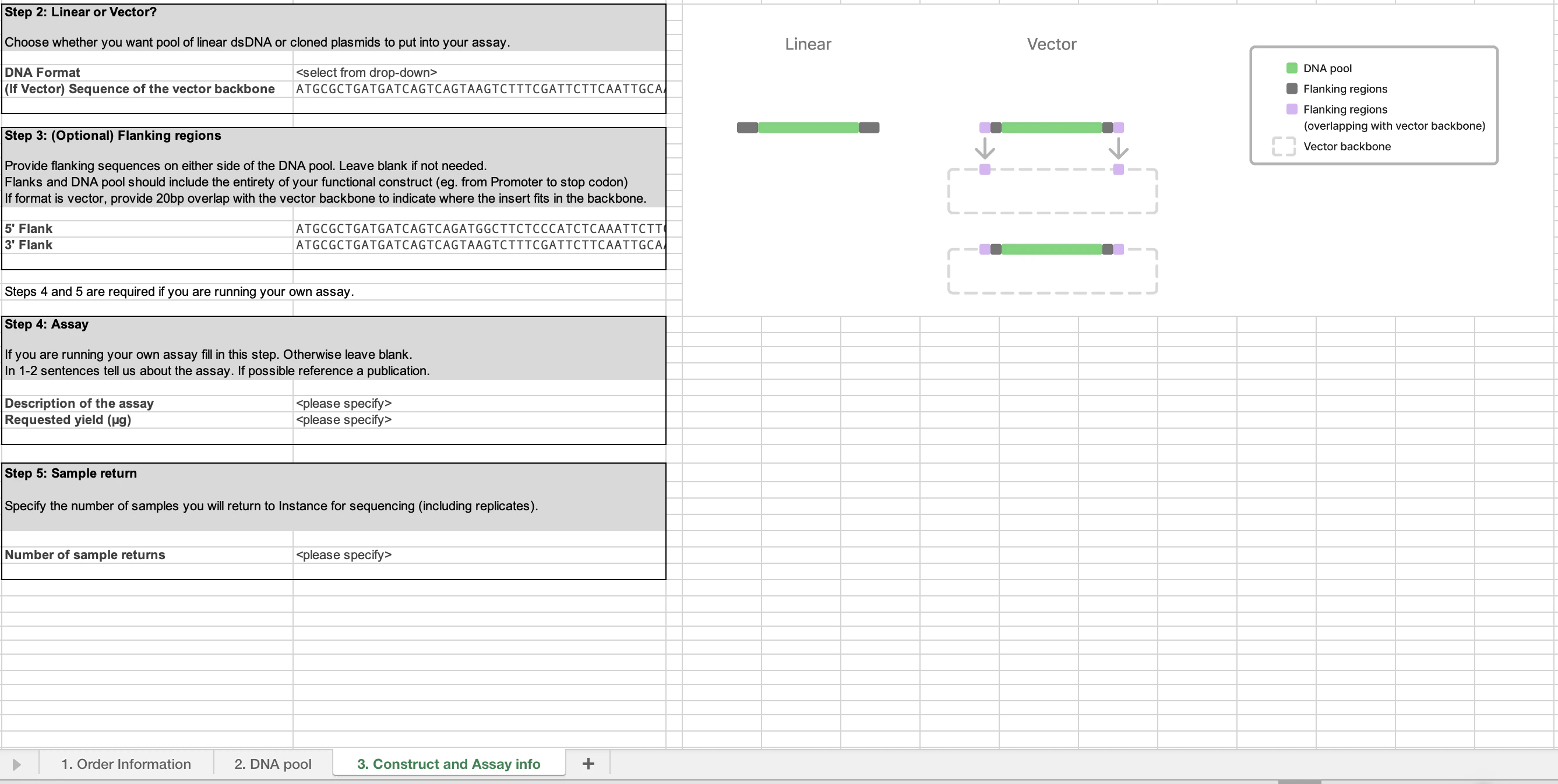
- The Construct and Assay info tab (3) collects information on your assay format, vector information (if necessary) and how many samples you would like to return for data generation.
Step-by-step guide on how to fill out the order form
Fill out order information sheet
Customer information
Customer information
The first section of this sheet covers customer information. We request your company/institution that you are associated with. We then ask for information regarding the project this dataset will be associated with. This provides us the ability to provide analysis and insights from across individual datasets within a project. Finally, we request your preferred alias for this pool as it allows us to communicate with you regarding your order’s progress.
| Property | Instructions |
|---|---|
| Company/Institution | Fill in the company, institution, or lab for which this order is placed |
| Project name (optional) | Fill in a project name |
| Order Alias (optional) | Fill in your preferred alias for the order |
Shipping information
Shipping information
This section provides us information on where to send the DNA pools once synthesized as well as the plate for return of the samples and sequencing. We will also use the shipping contact to inform the customer on any order updates.
Shipping information only needs to be provided if you are running your own assay. If you are ordering end-to-end data generation you do not need to complete this section.
| Property | Instructions |
|---|---|
| Shipping contact | Contact for shipping DNA pool to |
| Shipping email address | Email address |
| Shipping address (full) | Full address of the entity we are shipping the DNA to |
| Shipping phone number | Phone number of the entity we are shipping DNA to |
Billing information
Billing information
Please fill in where you would like billing and invoicing statements to be delivered to.
| Property | Instructions |
|---|---|
| Billing contact | Billing contact |
| Billing email address | Billing email address |
| Billing address (full) | Billing address |
| Billing phone number | Billing phone number |
Provide your DNA pool details
This is where you enter the pool sequences you would like to test. They can be entered as amino acid sequences and these will be codon optimized, based on the host species you select. We also accept nucleotide sequences. We currently do not accept degenerate bases within the pool region. The maximum size accepted for each fragment is 1000 basepairs. 

Complete the Construct and Assay Info
Tab 3 contains the critical information for your specific assay. There are four steps to complete:
Linear or Vector?
Linear or Vector?
First, select if you would like your DNA pool provided as linear double-stranded (ds) DNA or as a cloned plasmid. If you would like the pool cloned into a plasmid, please specify the vector sequence. The pool will be cloned in between the last and first position of the vector sequence. The vector sequence should encompass the entirety of your construct, including promoters, stop codon etc.
Flanking Sequences
Flanking Sequences
If you have sequences that remain constant around the DNA pool sequence, you can put them here. This can be utilized with either dsDNA delivery or as a vector. If you are using a vector backbone, please paste 20 bp of the vector sequence upstream (5’) and downstream (3’) so we can confirm the location of the pool. 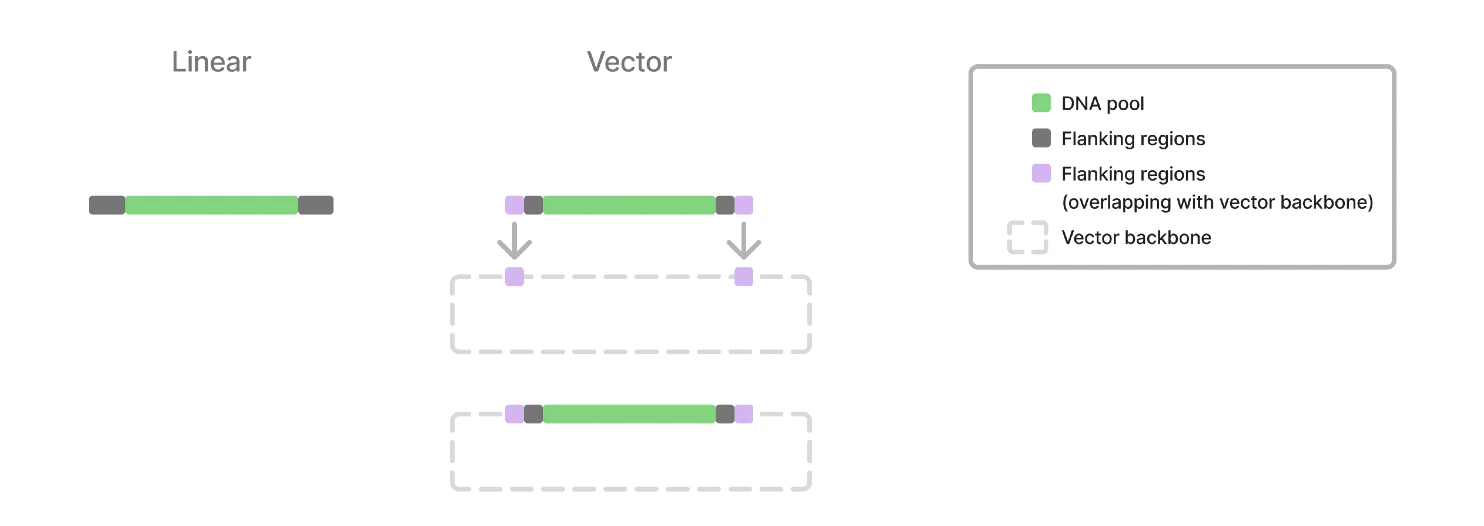

Assay
Assay
If you are running your own assay, please provide some context on the assay you are using this material for. Please specify how much material (in ug) you require. If you are utilizing end to end data generation, this section is not needed.
Assay Return
Assay Return
If you are running your own assay, please enumerate how many samples do you expect to generate. These will be returned to Instance for analysis and data generation. If you are utilizing end to end data generation, this section is not needed
Submit!
Please submit the order form via email to orders@instance.bio. Please ensure all three tabs have been completed.
Once we have received your sequences, we provide back a GenBank file to confirm the structure is as you intended.

